Quick Start
Input data format
Signal data of each trial should be stored as a 2d ndarray with shape (n_samples, n_channels), where each column represents data of one channel. If only one channel is presented, it can also be stored as a 1d ndarray with shape (n_samples,).
Timestamp data of each trial should be stored as a 1d ndarray with shape (n_samples,). If timestamp data is not provided, it will be generated automatically, starting from 0 and in increments of 1/hz, where hz is the sample rate.
Data of multiple trials should be organized in a list.
Demo code
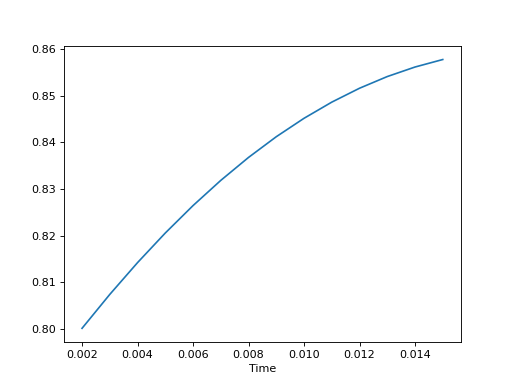
The demo code simply creates example signal data with shape (20,), assumes sample rate 1000 Hz, applies seven processing steps to the data using class EMGMeasurement, and plots the final result.
>>> import numpy as np
>>> import pyemgpipeline as pep
>>> data = np.array([20.3, 41.0, 53.9, 63.3, 39.5, 24.9, 26.1, 24.0, 44.1, 42.0,
... 37.4, 24.6, -21.8, -56.3, -48.1, -45.0, -29.1, -9.6, 5.3, 1.4])
>>> hz = 1000
>>> m = pep.wrappers.EMGMeasurement(data, hz=hz)
>>> m.apply_dc_offset_remover()
>>> m.apply_bandpass_filter()
>>> m.apply_full_wave_rectifier()
>>> m.apply_linear_envelope()
>>> m.apply_end_frame_cutter(n_end_frames=2)
>>> m.apply_amplitude_normalizer(max_amplitude=8.5)
>>> m.apply_segmenter(beg_ts=0, end_ts=0.015)
>>> m.plot()
(Source code, png, hires.png, pdf)